Design Protein Binders with NVIDIA's Proteina-Complexa
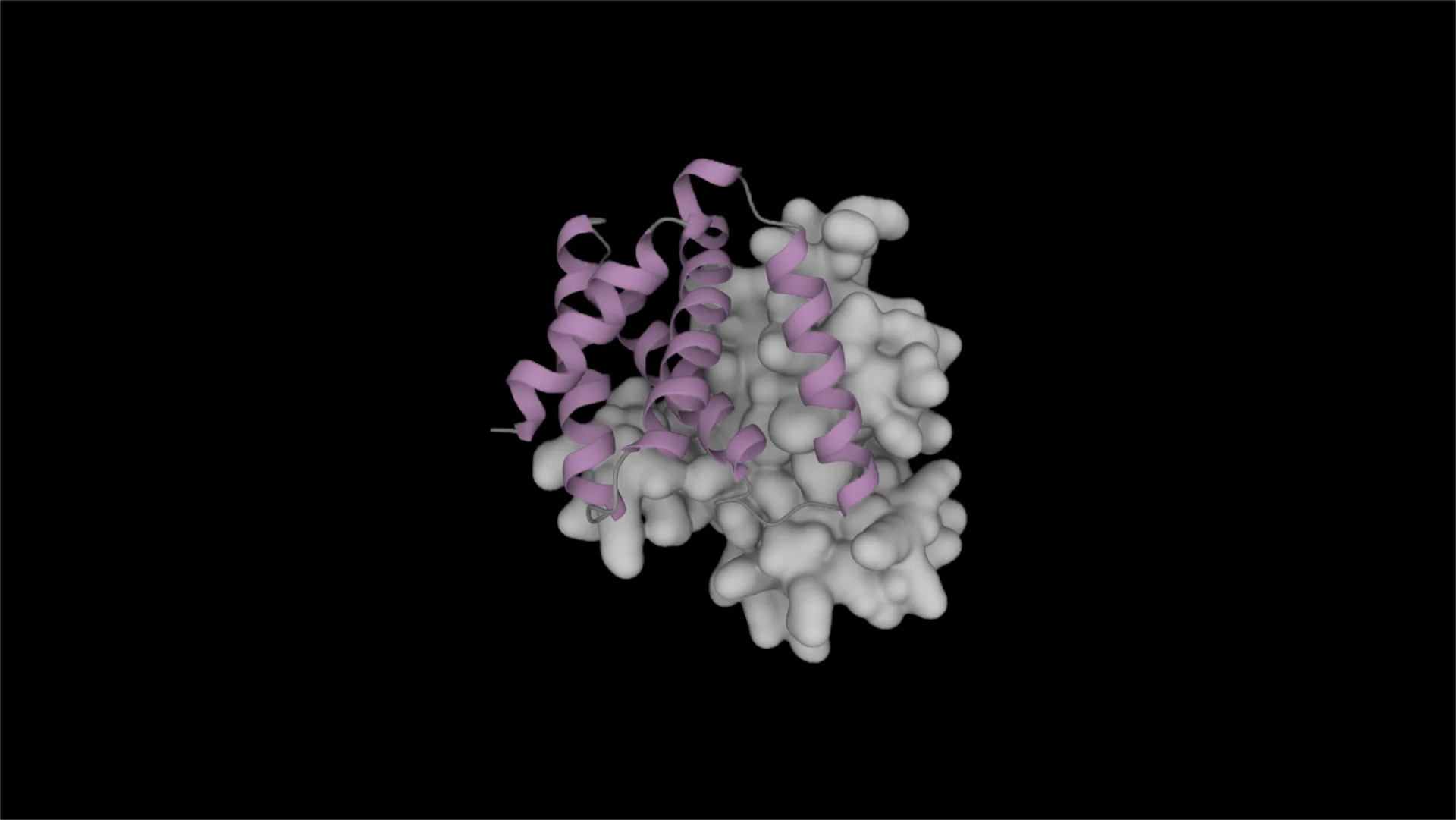
Developing new protein-based therapies and catalysts involves the challenging task of designing protein binders, or proteins that bind to a target protein or small molecule. The search space for possible amino acid sequence permutations and resulting 3D protein structures for a designed binder is vast, and achieving strong, specific binding requires careful optimization of the interactions between the protein binder and the target.
To address these challenges, NVIDIA has released Proteina-Complexa, a generative model that designs de novo protein binders and enzymes. In this article, we detail the key technologies behind Proteina-Complexa, explore primary use cases, and highlight the extensive experimental validation of generated protein binders. We also provide a step-by-step guide for using the command-line interface to generate your own binders.
The performance of Proteina-Complexa relies on three distinct technical components: the base generative model, the training datasets, and the integration of inference-time compute scaling. Proteina-Complexa generates protein binders through a partially latent flow-matching generation process combined with inference-time compute scaling to steer generation using reward functions. Built on top of the La-Proteina model, Proteina-Complexa uses a partially latent flow-matching framework to generate both fully atomistic binder structures and the corresponding amino acid sequence, called co-design.
This co-design approach enables reasoning at an atomistic level. By generating the amino acid sequence and the fully atomistic structure simultaneously, Proteina-Complexa ensures that the chemical identities and 3D geometry are tightly coupled. This integrated generation allows for the design of precise, high-affinity interfaces that are inherently optimized for folding and synthesis. Training a generative model for protein binder design requires a large amount of structural data on binders and their targets. Proteina-Complexa was trained on over 1 million curated, high-quality experimental and predicted structures from various databases.
Use cases for Proteina-Complexa include protein binders for protein targets and small molecule targets, as well as enzyme design. You can use Proteina-Complexa to design de novo protein binders against disease-relevant targets across indications including oncology, immunology, and neurology. This use case has been experimentally validated with collaborators from various institutions.
Simplifying Radar Data Processing with NVIDIA DRIVE for Autonomy
Maximize AI Factory Efficiency to Boost Revenue
Related articles

Creating a Pose2Sim Pipeline for Markerless 3D Kinematics
Learn how to create a Pose2Sim pipeline for markerless 3D kinematics.
Accelerating Protein Structure Prediction at Proteome-Scale
Exploring methods to accelerate protein structure prediction at proteome scale using AlphaFold and NVIDIA.

Anthropic keeps new AI model private due to cybersecurity vulnerabilities
Anthropic decided not to release the Claude Mythos Preview model due to cybersecurity vulnerabilities.